Making connections
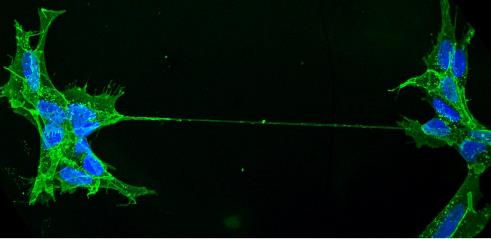
Mysterious nanotubes connecting distant cells have a key role in directly moving organelles – as well as viruses and disease-causing proteins – from one cell to another, write Diégo Cordero Cervantes and Chiara Zurzolo
The Biologist 66(2) p10-13
Everything we do, from vital functions such as breathing to more complex tasks like driving a car, is the result of a highly ordered level of communication between the cells that make up the tissues and organs in our bodies. This intercellular communication, also vital for the homeostatic orchestration of development and ageing, relies on the constant exchange of messages between our cells.
These messages are extraordinarily diverse in form and function, ranging from neurotransmitters and growth factors to hormones. Other modes of communication commonly used by our cells include gap junctions – nanoscopic channels formed by proteins that connect two adjacent cells and allow the passage of small molecules, ions and electrical signals; synapses – junctions that transmit chemical and electrical information between neurons; and exosomes – lipid vesicles released by cells that shuttle proteins and genetic information to both neighbouring and distant cells.
A more recently discovered mechanism for cell communication involves tunnelling nanotubes (TNTs)[1]. These are open-ended tubes that directly connect the cytoplasm of two (or more) distant cells.
For more than a decade, TNTs have been gaining international scientific attention for their use in the transfer of cellular contents, challenging the central dogma of cells – i.e that they are isolated entities. The combined data from various research groups around the globe, including ours, have shown that TNTs play a role in coordinating metabolism and signalling events in relation to cancer and infectious, genetic, metabolic and age-related diseases[2]. Both in vitro and in vivo studies have shown that TNTs enable the transport of entire organelles, bacteria, viruses, ions, genetic material and proteins between the cytoplasm of distant cells.
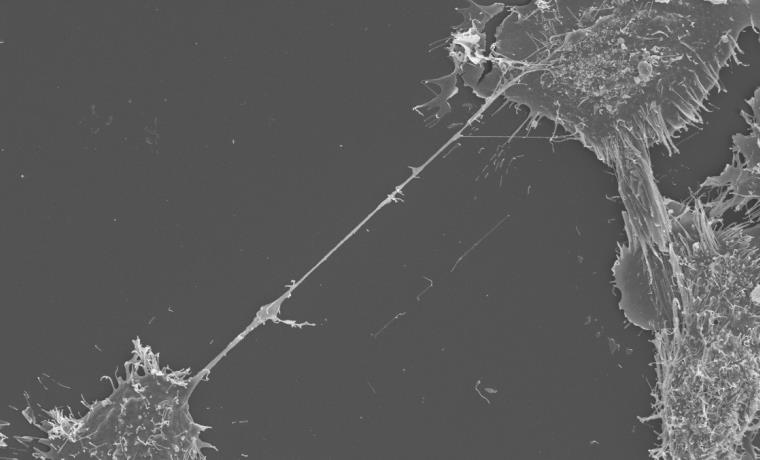 SEM image showing a TNT stretching between two neurons.
SEM image showing a TNT stretching between two neurons. Our laboratory has demonstrated that TNTs serve as the predominant pathway for the intercellular spreading of amyloids (misfolded aggregated proteins that lead to neuronal cell death) such as tau, alpha-synuclein and huntingtin. These misfolded proteins accumulate in the brain of patients with Alzheimer’s, Parkinson’s, and Huntington’s disease, respectively[3].
We discovered this behaviour when studying prions. Prions are transmissible pathogenic agents that are able to induce abnormal folding of normal cellular prion proteins, which are abundant in the brain. The abnormal folding of the prion proteins leads to brain damage and a family of progressive neurodegenerative disorders.
Our experiments demonstrated that prions are able to spread to other cells by hijacking TNTs[4]. The fact that amyloid proteins behave like prions, in their ability to induce misfolding of normal proteins, have led us to propose that TNTs may serve as one of the major avenues for the spreading of neurodegenerative diseases in the central nervous system. Therefore, TNTs represent a novel therapeutic target for minimising, or possibly even stopping, the progression of these incurable disorders.
So how do TNTs spread these harmful molecules? TNTs are highly dynamic tubes with lengths that range from several hundred nm up to 100µm. Their walls consist of a lipid bilayer contiguous with the plasma membrane of the cells it connects; the lumen of the tube establishes a direct connection between the cytoplasm of the linked cells. Inside these cylindrical structures lie cytoskeletal filaments called actin, which serve as tracks for the transport of ‘cargo’ powered by motor proteins. This dynamic membranous and cytoskeletal architecture facilitates the transport of cargo between cells. Why and how TNTs form is still not clear and is the subject of further investigation.
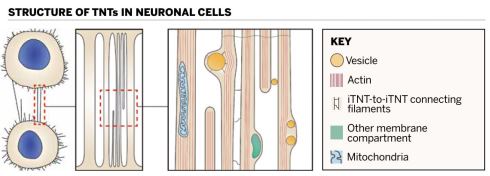 Filaments stretch down the length of individual tunnelling nanotubes (iTNTs) and connect the individual tubes to each other
Filaments stretch down the length of individual tunnelling nanotubes (iTNTs) and connect the individual tubes to each otherThus far, we have learned that TNT formation is largely generated by either the extension of a tubular membrane protrusion to another cell located at some distance, or between cells already in close proximity that then move apart and leave behind continuous intercellular nanotubes as the cells move in different directions.
Other cellular protrusions that resemble TNTs in appearance, but lack the ability to share cargoes between cells, are called filopodia – thin, actin-containing membranous spikes that act as antennae for cells to probe their surrounding environment. Previous work from our group has shown that filopodia and TNTs are regulated by the same actin cytoskeleton regulator protein complexes that surprisingly behave in an opposite manner, suggesting that these two structures have predetermined fates from the start[5]. Another pivotal piece of missing information is whether TNTs are open-ended at both cell-to-TNT contact sites, truly creating continuity between two cells. This is one of the most debated questions in the field of TNT research.
Taking a closer look at these two structures with enough magnification and detail would enable us to answer this question and better understand whether TNTs are unique structures, different from known cellular protrusions. However, carrying out experiments that can address these concerns is not as straightforward as it may seem. The fragile and transitory nature of TNTs makes them difficult to preserve for studies with scanning electron microscopy, which requires harsh sample preparations prior to high-resolution imaging.
To overcome these obstacles we combined a variety of confocal, 2D and 3D transmission electron microscopy techniques to characterise the anatomical architecture of TNTs in cells of different species. Using state-of-the-art microscopic approaches (e.g. correlative light- and cryo-electron tomography) in below-freezing temperatures (conditions optimal for the preservation of these delicate structures), we succeeded at contrasting TNTs’ structural identity with that of filopodia.
We found that most TNTs – seen in fluorescence microscopy as seemingly single connections – are in fact made up of two to 11 smaller, individual tunnelling nanotubes (iTNTs), all of which contain long, parallel actin filaments[].
Our electron tomogram images also suggest the existence of thin filaments and threads that connect iTNTs and presumably increase the mechanical stability of the overall bundle of tubes. Furthermore, we employed a powerful microscopic approach known as focused ion beam – SEM, or FIB-SEM, to produce 3D electron micrograph volumes with sufficient resolution to identify clearly that TNTs are ‘open’ at both ends and thus form a continuous cytoplasmic bridge between two cells. Finally, by using time-lapse imaging, we showed that the nanotubes transport mitochondria between cells.
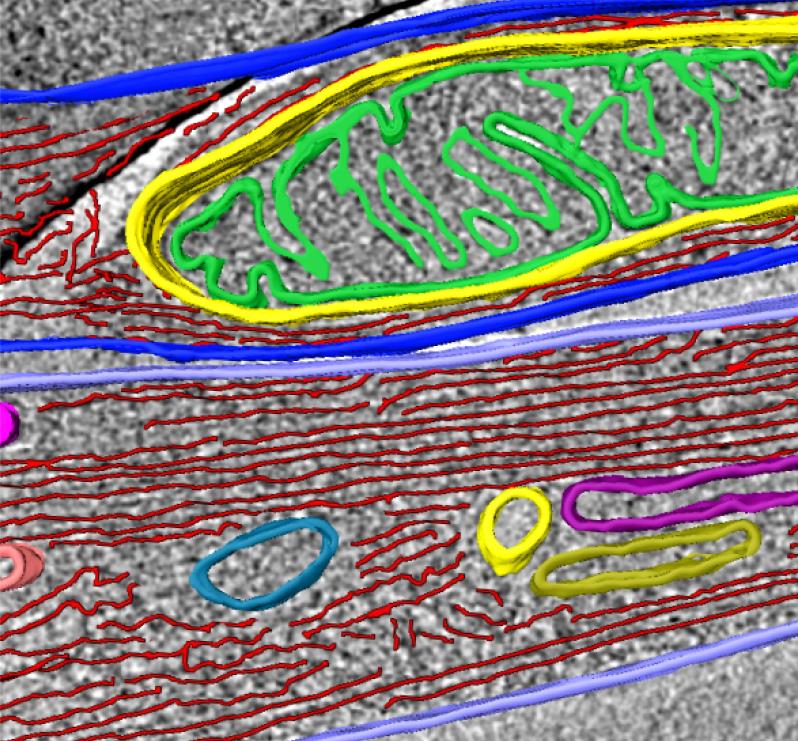 Mitochondria, actin, and membranous compartment revealed within an individual tunnelling nanotube (iTNT).
Mitochondria, actin, and membranous compartment revealed within an individual tunnelling nanotube (iTNT).Taken together, our discoveries challenge the dogma of cells as individual units by showing that cells can open up to neighbours and exchange materials and vital organelles without a membrane barrier. By applying an imaging workflow that overcomes previous limitations in tools used to study the anatomy of TNTs, we provided the first structural description of TNTs. Importantly, we demonstrated that TNTs are novel cellular organelles, with a defined structure, significantly different in morphology and function from known cellular protrusions such as filopodia.
This work is an essential step toward understanding TNT-mediated cell-to-cell communication, and lays the groundwork for investigations into the spreading of pathogens such as viruses, bacteria and misfolded proteins.
The description of the structure allows a better understanding of the mechanisms involved in their formation and provides detail on their function in transferring material directly between the cytosol of two connected cells. Essential questions about what signals trigger their formation, and how they establish specificity and selectivity, remain unanswered, however.
The answers to these and other fundamental questions merit tremendous scientific and clinical interest. Inhibiting the formation of TNTs may block or reduce the amplification of several diseases including HIV, Parkinson’s, Lewy bodies and multiple system atrophy, but first we must understand their function in non-pathologic states.
Fundamental research on how and when these structures form will be vital to any further work on the role of TNTs in disease.
1) Rustom, A. et al. Nanotubular highways for intercellular organelle transport. Science 303, 1007–1010 (2004).
2) Ariazi, J. et al. Tunneling nanotubes and gap junctions: their role in long-range intercellular communication during Development, Health, and Disease Conditions. Front. Mol. Neurosci. 10, 333 (2017).
3) Abounit, S. et al. Tunneling nanotubes: A possible highway in the spreading of tau and other prion-like proteins in neurodegenerative diseases. Prion 10(5),
344–351 (2016).
4) Gousset, K. et al. Prions hijack tunneling nanotubes for intercellular spread. Nat. Cell Biol. 11(3), 328–336 (2009).
5) Delage, E. et al. Differential identity of filopodia and tunneling nanotubes revealed by the opposite functions of actin regulatory complexes. Sci. Rep. 6, 39632 (2016).
6) Sartori-Rupp, A. et al. Correlative cryo-electron microscopy reveals the structure of TNTs in neuronal cells. Nature Communications 10(1), 342 (2019).
Diégo Cordero Cervantes is a graduate student at the Unit of Membrane Traffic and Pathogenesis at the Pasteur Institute, France. Chiara Zurzolo is head of the Unit of Membrane Traffic and Pathogenesis at the Pasteur Institute, France.


