'There’s still something new every day'

To celebrate 100 years since the formation of the Genetics Society, Alison Woollard talks to geneticist Jonathan Hodgkin about a centenary of remarkable progress and what they both owe to mutant worms
The Biologist 66(6) p14-17
Jonathan Hodgkin FRS is emeritus professor of genetics at the University of Oxford and a former president of the Genetics Society. In the 1970s he was a student of Sydney Brenner, the first researcher to suggest the nematode worm Caenorhabditis elegans could be used to study fundamental biological processes. C. elegans went on to become a key model organism in genetics, development and neuroscience and has been central to Hodgkin’s career ever since. It was the first multicellular organism to have its whole genome sequenced and remains the only organism to have had its entire ‘connectome’ (neural wiring) mapped.
Hodgkin’s early work at the Laboratory of Molecular Biology in Cambridge helped elucidate the complex genetic pathways that determine whether individual worms become male or hermaphrodite, but in his long career he has explored many other developmental pathways and the genetic basis of behaviour. More recently Hodgkin has focused on the nematode’s innate immunity to attack by various bacteria, which has helped identify antibacterial factors that could be candidates for new antibiotics.
Hodgkin was elected a Fellow of the Royal Society (FRS) in 1990 and received the Genetics Society Medal in 2011. He is the son of Nobel Prize-winning biologist Alan Lloyd Hodgkin.
AW: Jonathan, it’s really great to be talking to you as we celebrate a century of genetics. What got you into it?
JH: Largely brilliant teaching at university. I remember reading a genetics textbook in sixth form and not understanding a word of it. A couple of years later it all made sense thanks to, among others, a particularly good fungal geneticist called Brian Cox. No, not that one.
Then, as I became interested in developmental biology it was obvious that genetics was going to be the most powerful tool to work out what exactly was going on with development.
Tell us how you got started looking at C. elegans.
It was because of Sydney Brenner, who had realised that you could use this organism in enormously powerful ways. I heard a lecture from him in my final year of university and was totally inspired by that. When I graduated I persuaded him to take me on as his graduate student. That was a very, very good decision.
I mean, he was a great believer in choosing the right organism for what he worked with. He went from phages and bacteria to C. elegans, and then got involved with various genome projects.
He was interested in fish too, wasn’t he?
He liked fish, yes, among other things. His first paper was on the chromosomes of elephant shrews in South Africa. Later on he rather neatly did the complete genome of a weird creature called the elephant shark.
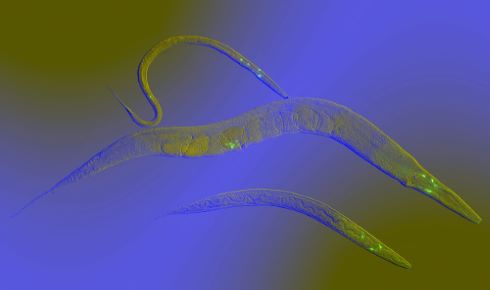
Switching sex C.elegans is unusual among nematodes in that individuals are either males (XX) or hermaphrodites (XO) that can self-fertilise. Males can inseminate hermaphrodites, which will preferentially use male sperm rather than their own male gametes. Hodgkin’s work unpicked the regulatory cascade of genes that control whether a worm develops male or hermaphrodite characteristics. He identified that mutation of the tra-1 gene can completely switch an individual’s sexual phenotype.
Did anyone think you were a bit mad to follow Sydney’s call to study C. elegans?
Some people were very sceptical of the whole C. elegans project. Other people had faith in Sydney – they could see that this was basically a genius who had settled on a terrific system and it was going to go places. He also inspired a lot of other people to join him, so in the end it wasn’t just about Sydney, it was all the other people that came in.
Are there other model organisms that you would really like to have seen succeed that just didn’t make it?
The one that I think is particularly sad is this amoeba called Naegleria, which we now know from its genome is a terrific basal eukaryote and does all sorts of interesting things, as amoeba go. It has features superior to Dictyostelium.
The trouble with Naegleria is that it can kill you by eating your brain. So people never really took to it…
It’s the centenary of the Genetics Society, so we’re celebrating 100 years of genetics. In the whole field of genetics what discoveries stand out for you over the last hundred years?
Oh, any number of things. The establishment of chromosome theory and heredity; what Thomas Hunt Morgan did putting the genetics together with the cells. It’s a pretty heroic story to go through. Then people start being able to do molecular genetics on fungi, like [George] Beadle and [Edward] Tatum with their idea of ‘one gene for one enzyme’.
Then you move into a whole other layer with the DNA double helix and genetic code, and understanding the basics of transcription and translation.
Then other important things were happening at the same time – for example, the idea that the genome is malleable and has mobile elements, which is down to Barbara McClintock. All of that is amazing. And it relied on model organisms – Morgan with flies, McClintock working with maize, Beadle and Tatum with the mold Neurospora.
Subsequently, all the people working on the phage and with Escherichia coli worked out molecular genetics – the ability to clone and amplify DNA was fantastically important, because that really allowed us to get into the biochemistry of it all. And that was all dependent on the ability to sequence DNA, which was down to Fred Sanger and others. This just got more and more efficient and then all of it was succeeded into the whole genetics revolution, which was and continues to be quite extraordinary, and changes everything.
What do you think is going to be the most exciting development over the next 10 to 20 years?
I think a lot is going to come out of the large-scale genome analyses people are doing. At the moment everybody is basically drowning in data and you have to say that a lot of what has come out has been disappointing. But people should not despair.
Do you think that’s just the bottom of an exponential curve that will come?
Yes, I think it’s going to come. I think there is a wood and trees problem, particularly with phenomena that we can see are very, very multigenic. However, I think people will get through that and we will see that you can understand what’s going on by focusing on a few of the key players.
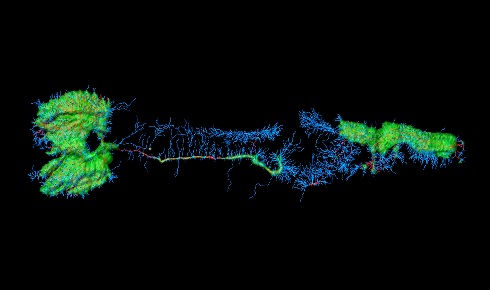 Why worms? All laboratory strains of C. elegans are thought to be derived from one individual isolated from soil in Bristol in the 1950s. These tiny worms, around 1mm in length, replicate roughly every three days and are transparent, making them ideal for studying genetic and developmental changes. They are easy to grow, with many hundreds fitting on a single agar plate, and remain viable after being frozen and thawed. With just 1,000 cells in each adult worm (not including germ cells), researchers are able to understand and follow the developmental fate of every individual cell in the organism.
Why worms? All laboratory strains of C. elegans are thought to be derived from one individual isolated from soil in Bristol in the 1950s. These tiny worms, around 1mm in length, replicate roughly every three days and are transparent, making them ideal for studying genetic and developmental changes. They are easy to grow, with many hundreds fitting on a single agar plate, and remain viable after being frozen and thawed. With just 1,000 cells in each adult worm (not including germ cells), researchers are able to understand and follow the developmental fate of every individual cell in the organism. What do you think about the capacity we now have to interfere with our genomes and change them? Do you think that’s an exciting opportunity? Is there anything that worries you about that?
I think that it’s tremendously exciting. I think that it’s also tremendously worrying and that a lot of this is going to happen willy-nilly, and we just have to live with this. The human race is not a static entity. This technology is not going to go away, either.
Coupled with what we were just saying about complexity, this means a lot of these genetic changes that people hope will make their baby wonderful – it’s going to be totally different. Not totally unpredictable, but there are going to be huge numbers of unpredictable consequences.
People seem to often think of genetics as being deterministic and I think that’s a big mistake because it’s probabilistic. I think that’s quite a challenge to discuss with people who don’t really know about genetic principles.
It certainly is. Putting things to lay people is always difficult whatever area you’re in, but the public and politicians are not good about thinking about probability, nor risks. We have obviously seen this with climate change and other environmental issues – you’re dealing with chances, you’re not dealing with certainties.
You’ve spent most of your career staring down a microscope at these tiny C. elegans worms. You can hear in the way you talk about them that you still have a huge passion. What is it about these worms that fosters that?
Simply the fact that there’s something new every day after all these years. But also because the worm grows so fast – three days’ generation time – you can think of something, experiment and get a result in a few days. It’s the practicality of the worm that has a big appeal: the economy and the lack of ethical issues.
And you can see so much because they are transparent…
Yes, they are kind of fun to look at. We now have this enormous body of knowledge on them. There’s also a huge amount we don’t understand. It’s just very satisfying, going on doing experiments, confirming things.
If somebody gave you £10m to work on something new or something that you maybe had not had time to pursue in the past, what do you think you might do?
There are really ambitious things that you can think about pursuing – doing a very large amount of genome editing to try to make a ‘minimal worm’ with as few genes as possible, for example. People are doing some of that already and it is a fascinating way to go.
You could also try to do more in the way of humanising the worm. Sticking in human genes or human pathways so that you could then study them in this system.
One simple thing that is bugging me at the moment is almost everything we do is looking at worms behaving in a two-dimensional sheet, whereas that’s not their real environment. They live in a three-dimensional world. There are whole bits of their nervous system and sensory organs and we don’t know what they are for.
All these scientists have been beavering away for years on this tiny worm and there’s still so much that we don’t know. If you widen that view out to the rest of the other model organisms, and then of course all the other organisms on Earth, and then humans, you can see that biology has a really long way to go.
It does. One of the conclusions from that is that the only organism we are going to achieve any kind of complete understanding of is probably humans.
What would you do if you were starting out again?
One thing would be to work on something with practical utility, such as tuberculosis. I think diseases are wonderful ways into biology, but they are also important.
The other would be to get back to the large-scale problem that really got me to this business in the first place, which is how you genetically encode complex behaviour.
The dog genome projects are great because dogs are so various in their behaviour and cognition. You have the genetics, you have the animals and you have enthusiasm. I think that it’s actually going to tell us some significant things.
And they’d be such wonderful creatures to look at every day as well…
Yes, exactly!


